CAP TODAY and the Association for Molecular Pathology have teamed up to bring molecular case reports to CAP TODAY readers. AMP members write the reports using clinical cases from their own practices that show molecular testing’s important role in diagnosis, prognosis, and treatment. The following report comes from Beth Israel Deaconess Medical Center and Harvard Medical School. If you would like to submit a case report, please send an email to the AMP at amp@amp.org. For more information about the AMP and all previously published case reports, visit www.amp.org.
Loren Joseph, MD
Derek Jones, MD
October 2020—After 20 years of CAP advocacy, synoptic reporting in surgical pathology is ubiquitous. This came about in part by fiat and in part by all parties agreeing on the importance of standardization for patient care. The merits of some elements remain controversial.1 Molecular pathology, a newer discipline, does not offer the scope for creative writing once available in surgical pathology. Molecular pathology reporting is not fully synoptic, but CAP checklists already mandate several critical elements:
Human sequence variants must be reported using HGVS nomenclature and include the HUGO Gene Nomenclature Committee (HGNC) gene name, and a standard versioned reference identifier to the transcript/protein (eg, REFSeq Accession Number, Ensembl Transcript/Protein ID, or CCDS ID) that allows unambiguous mapping of the variant. The reference genome (if applicable) assembly and version number used for alignment and variant calling must be reported. Variant chromosomal position (ie, genomic coordinate) should be reported. (Molecular pathology checklist, Aug. 22, 2018, MOL.36155 Sequence Variants.)
This still leaves several elements open to laboratory discretion, such as variant allele fraction (VAF) and gene copy number.
Despite payers’ reluctant recognition of the utility of next-generation sequencing panels, the scope and frequency of testing continue to expand. It is now common to have serial testing for hematopoietic malignancy; this can reveal interlaboratory variability.
Case. The patient was a 72-year-old female referred from an outside hospital, where she had presented with a complaint of three months of increasing fatigue. Workup showed mild pancytopenia: WBC 3.6 × 103/μL (ref. 4.0–10.0), Hgb 10.3 g/dL (ref. 11.2–15.7), Hct 33.6 percent (ref. 34–45), MCV 70 fL (ref. 82–98 fL), platelet 148 × 103/μL (ref. 158–400), neutrophils 35.9 percent (ref. 34–71), lymphocytes 40.4 percent (ref. 19–53), monocytes 19.8 percent (ref. 5–13), eosinophils 2.2 percent (ref. 1–7), basophils 0.3 percent (ref. 0–1).
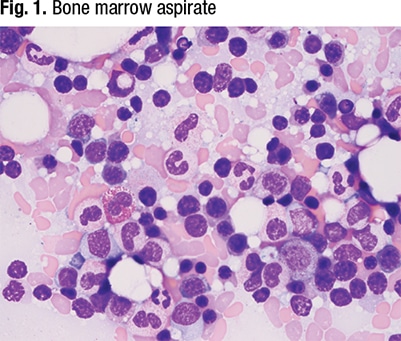
Her prior medical history was notable only for mild beta-thalassemia. She had no family history of malignancy. A bone marrow aspirate (Fig. 1) was sent by the outside hospital for a myeloid gene panel at Lab No. 1.
The report noted only a variant in the TET2 gene, c.4393 C>T (p.R1465*) (human genome build hg19), with a VAF of 40 percent and 2,000× coverage at that position. This notation indicates that at nucleotide position 4393, a cytosine has changed to a thymine, and in the corresponding protein sequence the arginine, amino acid 1465, has been changed to a stop codon, a nonsense variant, truncating the protein product. The VAF of 40 percent implies that the noted TET2 variant may be present in as many as 80 percent of nucleated cells in the sample, if diploidy is assumed.
 The patient was referred to our hospital for evaluation of a myeloid disorder. The patient had not received any treatment. Our review of the outside bone marrow biopsy showed normal trilineage hematopoiesis without dysplasia. A peripheral blood was sent for a myeloid panel at Lab No. 2. No variants were found. In the absence of a variant, Lab No. 2 does not indicate the coverage for each gene, but the report gives the average coverage across the panel, for this sample, as >1,000× (range 500–50,000). Lab No. 1 and No. 2 assays have similar stated general sensitivity, five percent VAF. The treating physician, concerned about myelodysplastic syndromes and clonal hematopoiesis of indeterminate potential, wants to know which laboratory to trust.
The patient was referred to our hospital for evaluation of a myeloid disorder. The patient had not received any treatment. Our review of the outside bone marrow biopsy showed normal trilineage hematopoiesis without dysplasia. A peripheral blood was sent for a myeloid panel at Lab No. 2. No variants were found. In the absence of a variant, Lab No. 2 does not indicate the coverage for each gene, but the report gives the average coverage across the panel, for this sample, as >1,000× (range 500–50,000). Lab No. 1 and No. 2 assays have similar stated general sensitivity, five percent VAF. The treating physician, concerned about myelodysplastic syndromes and clonal hematopoiesis of indeterminate potential, wants to know which laboratory to trust.
Resolution. In the COSMIC database (cancer.sanger.ac.uk/cosmic, accessed March 5, 2019), there is data from some portion of the TET2 gene from more than 65,000 samples. This particular nonsense TET2 variant is reported in 43 cases, all hematological malignancies, mainly myeloid, but including 13 cases of non-Hodgkin’s lymphoma, a lymphoid disorder. The variant is predicted to be pathogenic based on a protein modeling algorithm using a version of the FATHMM (Functional Analysis Through Hidden Markov Models) program at the website. The high VAF raises the possibility this variant is in the germline, but the SNP database at the NCBI (www.ncbi.nlm.nih.gov/snp) does not contain this variant. TCGA data for myeloid disorders (www.cBioPortal.org, accessed March 5, 2019) showed no evidence of this variant in 600 cases of AML, which were analyzed by whole exome or whole genome sequencing.
Pathogenic TET2 variants are found with increasing frequency with patient age and can reflect clonal expansions of abnormal cells with increased but indeterminate malignant potential. Still, the presence of a pathogenic TET2 variant alone is insufficient for a diagnosis of a myelodysplastic syndrome without morphological evidence. The bone marrow biopsy showed trilineage development with no evidence of dysplasia. There is increasing evidence that the presence of such variants does raise the risk of development of a myeloid leukemia and might be a corollary of a process that explains the pancytopenia.2 So far the case is not puzzling, although it is not clear that the pancytopenia has to be attributed to the presence of this variant.
Although peripheral blood does not always closely parallel the bone marrow, given the high VAF in the first marrow, one would expect at least a low level in the blood. The lack of a variant in the second marrow might be explained by patchy involvement, but combining two ad hoc explanations is unsatisfying. Scrutiny of the reports, however, revealed bioinformatic incommensurability.
As for many genes, there are multiple transcript isoforms for TET2 (Table 1).3
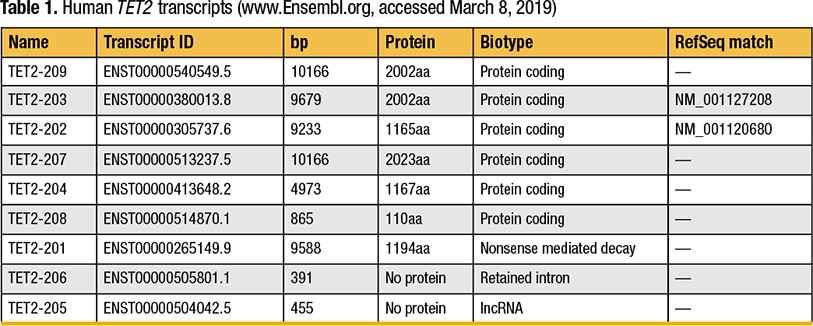
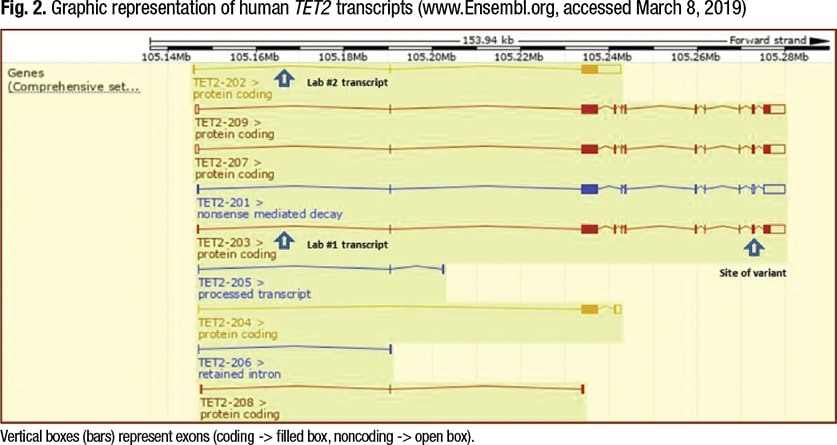
The general information in the report for Lab No. 1 awkwardly states, “All genes covered are not all sequenced in their entirety,” but usefully lists the reference transcript for TET2 as NM_001127208.2 (TET2-203 in Fig. 2, page 70). General information in the report for Lab No. 2 states that for all genes, “all protein coding regions plus splice sites” are covered by amplicons, and cites the reference transcript, NM_017628.4 (TET2-202 in Fig. 2). Lab No. 2 chose this transcript because it was listed as the reference transcript at the HGNC website when the assay was developed.
Transcript NM_001127208.2 encodes a longer predicted protein than does the transcript NM_017628.4 (Fig. 2).
The TET2 R1465* variant was not detected by Lab No. 2 because that region is not covered in its assay. Whether or not Lab No. 2 should target the larger transcript is a separate question. Short of exome or whole genome sequencing, every lab must balance multiple considerations in determining which regions to target. Lab No. 1 reported the variant only in terms of the reference transcript but not in terms of the reference genome. Lab No. 2 routinely reports a variant with respect to both the reference transcript and genomic reference coordinates, but since Lab No. 2 did not identify a variant, no coordinates were available for comparison. In this case, interpretation by the busy clinician would require looking up both transcripts and locating and comparing the variant(s) and/or transcripts, prompting consultation. In this case, the clinician accepted the result showing a TET2 variant and continues to monitor the patient without treatment. Alternative causes for the fatigue and pancytopenia have not been identified. The patient remains stable.
This case illustrates the difficulty in comparing results when busy clinical “consumers” understandably emphasize the “topline.” This can be especially problematic when a result is negative and no variant is reported. Reporting of the gene CEBPA illustrates this. It is an important gene in myeloid disorders, included in most hematologic malignancy panels, and known to be difficult to amplify and sequence because of its high content of GC nucleotides. A third reference laboratory we have used in the past never reported a CEBPA variant as topline even though the gene is “on” the panel. Review of the detailed coverage table always showed the CEBPA amplicons as absent.
The importance of formatting results for the user has received little attention in the pathology literature in general, even less for NGS.4-6 The presentation of data-dense next-generation sequencing results might defy even user-friendly interfaces for busy clinicians and can benefit from molecular pathology consultation, such as at tumor boards. Reporting both the reference transcript ID and the genomic coordinates as directed by the CAP checklists is a good place to start. Expecting all labs to use the same reference transcript, even if the entire transcript is not “covered” to the same extent in the assay, would further simplify comparisons. If a consensus committee were to recommend reference transcripts, it would require rewriting portions of bioinformatic pipelines, which is not trivial but would not require altering assays. The extent of variation in reference transcripts used in actual practice for a given gene has not been described to our knowledge, so the extent of this problem remains to be determined.
- Renshaw AA, Mena-Allauca M, Gould EW, Sirintrapun JS. Synoptic reporting: evidence-based review and future directions. JCO Clinical Cancer Inform. 2018;2:1–9.
- Steensma DP. How I use molecular genetic tests to evaluate patients who have or may have myelodysplastic syndromes. Blood. 2018;132(16):1657–1663.
- Wong ACH, Rasko JEJ, Wong JJ. We skip to work: alternative splicing in normal and malignant myelopoiesis. Leukemia. 2018;
32(5):1081–1093. - Powsner SM. Clinicians are from Mars and pathologists are from Venus. Arch Pathol Lab Med. 2000;124(7):1040–1046.
- Valenstein PN. Formatting pathology reports: applying four design principles to improve communication and patient safety. Arch Pathol Lab Med. 2008;132(1):84–94.
- Makhnoon S, Shirts BH, Bowen DJ, Fullerton SM. Hereditary cancer gene panel test reports: wide heterogeneity suggests need for standardization. Genet Med. 2018;20(11):1438–1445.
Dr. Joseph is assistant professor, Department of Pathology, Harvard Medical School, and director of the molecular diagnostics laboratory, Beth Israel Deaconess Medical Center, Boston. Dr. Jones, formerly of Harvard Medical School and Beth Israel Deaconess Medical Center, is a uropathology fellow at NYU Langone Medical Center, New York, NY.
[divider]
Test yourself
[tabs type=”horizontal”][tabs_head][tab_title]Here are three questions taken from the case report. [/tab_title][tab_title]Answers[/tab_title][/tabs_head][tab]
1. Which of the following elements of NGS reports for tumors is not required by the CAP checklist?
a. The official gene name.
b. The reference genome assembly (for human genomics).
c. The gene copy number (when variation is found).
d. A versioned reference to the transcript/protein.
e. Details of the software used for sequence alignment.
2. Which of the following statements is true about the existence of different transcripts for a given gene?
a. A given tissue will show only one kind of transcript.
b. A multiplicity of transcripts results only when there are mutations in splicing factor genes like SF3B1.
c. A point mutation, which changes an encoded amino acid (substitution mutation), will have the same designation when using proper HUGO nomenclature no matter which transcript is used as a reference.
d. A point mutation, which changes an encoded amino acid (substitution mutation), will have the same nucleotide designation by the genomic reference coordinate no matter which transcript is used as a reference for the amino acid encoding.
e. A point mutation, which changes an encoded amino acid (substitution mutation), will have the same nucleotide designation no matter which genomic reference build (version) is used.
3. A 65-year-old patient with persisting mild anemia refuses a bone marrow biopsy. The peripheral smear shows no dysplasia. His blood was sent for a myeloid cancer panel. The only finding is a TET2 likely pathogenic variant with a VAF of 30 percent. The patient is best described as having what?
a. A germline variant.
b. Clonal hematopoiesis of indeterminate potential.
c. Myelodysplastic syndrome.
d. Myeloproliferative disorder.
e. Acute myelogenous leukemia.
[/tab]
[tab]
Answers are online now at www.amp.org/casereports and will be published next month in CAP TODAY.
[/tab]
[/tabs]

