CAP TODAY and the Association for Molecular Pathology have teamed up to bring molecular case reports to CAP TODAY readers. AMP members write the reports using clinical cases from their own practices that show molecular testing’s important role in diagnosis, prognosis, and treatment. The following report comes from the Hospital of the University of Pennsylvania. If you would like to submit a case report, please send an email to the AMP at amp@amp.org. For more information about the AMP and all previously published case reports, visit www.amp.org.
Neha Gupta, MD; Selina Luger, MD; Dale M. Frank, MD; Jacquelyn J. Roth, PhD; Salvatore F. Priore, MD, PhD
February 2022—Myeloproliferative neoplasms (MPNs) are a group of clonal hematopoietic stem cell disorders characterized by increased proliferation of myeloid cells of one or more lineage. The most common MPNs include chronic myeloid leukemia (CML), polycythemia vera (PV), essential thrombocythemia (ET), and primary myelofibrosis (PMF). CML is defined by the BCR-ABL1 fusion, which typically results from the t(9;22)(q34;q11.2) rearrangement.1 JAK2, CALR, and MPL alterations are common in non-CML MPNs (the “classical MPNs”), though not exclusive to any one entity. The most common variant in PV is JAK2 p.V617F, a pathogenic gain-of-function variant observed in more than 95 percent of cases. Approximately three percent of JAK2 p.V617F-negative PV cases will show a JAK2 exon 12 variant. PMF and ET show a similar frequency of genetic alterations, with 50 to 60 percent harboring JAK2 p.V617F, 30 percent harboring CALR variants, and five to eight percent of PMF and three percent of ET displaying variants in the MPL gene. The remaining 10 to 15 percent of PMF and ET are considered triple negative and may contain an assortment of rare genomic variants.1,2
A heterogeneous collection of rare myeloid neoplasms with the “ungainly designation of myeloid/lymphoid neoplasms with eosinophilia (MLN-Eo) and rearrangement of PDGFRA, PDGFRB, or FGFR1, or with PCM1-JAK2” (WHO 2017),3 lacks the genetic findings that characterize CML and the other classical MPNs. Instead, these neoplasms harbor characteristic recurrent gene rearrangements involving the abovementioned tyrosine kinase genes. Clinically, constitutive kinase activation drives eosinophilia, which has been a useful organizing principle for this group of rare disorders. However, it has become clear that many patients present without significant eosinophilia and with clinical and pathologic findings that resemble a variety of hematologic neoplasms ranging from myelodysplastic syndromes to CML and the classical MPNs. Therefore, these diseases should be considered as part of the differential diagnosis for MPNs without an identified causative genomic variant.3,4 We present a case that highlights the important roles molecular diagnostics and variant classification play in establishing a correct diagnosis and optimal treatment plan for patients who present with features consistent with an MPN.
Case presentation. A 41-year-old female with a past medical history of surgically managed cervical cancer presented at an outside hospital with cough, dyspnea, congestion, malaise, abdominal pain, sweats, and myalgias. MRI of the abdomen revealed hepatosplenomegaly, liver cysts, and mild retroperitoneal lymphadenopathy. A complete blood count showed a mild leukocytosis, a myeloid left-shift, and nucleated red blood cells. Hemoglobin and platelets were within normal limits.
The patient was referred to a hematologist for further evaluation. A repeat CBC confirmed these abnormalities. Outside bone marrow studies showed morphologic abnormalities (marked hypercellularity, mild megakaryocytic atypia, and grade one reticulin fibrosis) diagnostic of an MPN and favoring primary myelofibrosis. Cytogenetic studies performed on a hemodilute bone marrow aspirate were non-diagnostic; fluorescent in situ hybridization studies were negative for a BCR-ABL1 rearrangement. Molecular studies (real-time PCR followed by melting curve analysis) performed on the bone marrow aspirate were reported as “positive for CALR exon 9 insertion/deletion mutation.” The resulting cDNA and protein changes were reported as “c.1142_1144delAGG” and “p.E381fs,” with an estimated allele fraction (VAF) of 50 percent. All CALR nomenclature in this case report is consistent with reference sequence NM_004343.3. The targeted molecular studies were negative for JAK2 p.V617F, JAK2 exon 12, JAK2 exon 13 (p.G5712S), and MPL W515L/K variants.
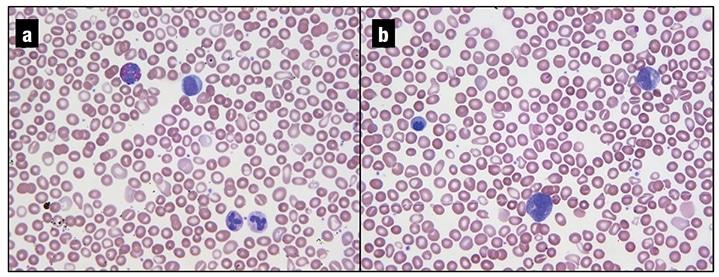
The patient was then referred to our institution for a second opinion. Review of a peripheral smear showed a leukoerythroblastic picture with teardrop forms (Fig. 1). CBC findings were similar to those noted previously: WBC 12.9 THO/µL (neutrophils 61.4 percent, bands one percent, metamyelocytes one percent, myelocytes one percent, promyelocytes two percent, lymphocytes 16.8 percent, monocytes 11.9 percent, eosinophils 2.9 percent, basophils one percent, blasts one percent), Hgb 13.3 g/dL, MCV 96 fl, platelets 176 THO/µL. Review of the outside hospital’s bone marrow study confirmed the diagnosis of myelofibrosis and noted mild eosinophilia (eight percent) on the aspirate smear.
Massive parallel sequencing (MPS) studies were performed in our institution’s CAP/CLIA-certified laboratories on a peripheral blood specimen collected one month after the initial outside hospital’s bone marrow biopsy. Our MPS assay is a hybrid-capture–based comprehensive genomic panel designed to detect single nucleotide variants, insertions and deletions, and select copy number gains in 116 genes with therapeutic, diagnostic, and/or prognostic importance for hematological malignancies. Data analysis was performed using a custom bioinformatics pipeline. All variants were annotated with reference to the hg38 genome build. The patient’s sample was positive for one missense variant of uncertain significance in the NFKBIE gene, c.266C>T, p.A89V, with a VAF of 51 percent (NM_004556.2). Sequencing did not detect pathogenic CALR exon 9 deletions, JAK2 exon 12 deletions, JAK2 p.V617F, or MPL codon p.W515 variants.
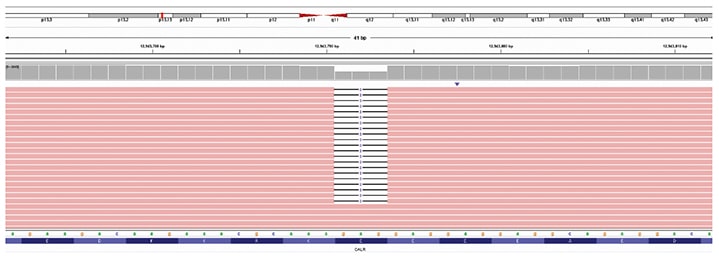

Given the apparent discrepancy between the initial molecular study, which reported a CALR exon 9 frameshift variant, and the in-house MPS study, which did not, the molecular findings of the outside hospital were reviewed. This revealed an incorrect predicted protein annotation for the reported c.1142_1144AGG variant, which is an in-frame deletion and not a frameshift variant. When we filtered our MPS data for in-frame deletions in CALR, the same c.1142_1144delAGG variant (more concisely described as c.1142_1144del) was identified (Fig. 2). As this variant was internally classified as a benign change, it was not flagged for review or reporting during our variant review workflow.
Given essentially negative MPS results, the referring oncologist ordered additional studies, all of which were performed outside our institution. A rearrangement of JAK2 (9p24.1) was detected in 52 percent of cells (156/300) using an investigational-use-only dual-color breakapart probe set (CytoCell). This result was confirmed by conventional cytogenetic analysis with a 46,XX,t(8;9)(p22;p24)[10]/46,XX[10] karyotype. An investigational-use-only MPS panel (Archer FusionPlex) also detected an in-frame fusion between PCM1 exon 36 (NM_006197) and JAK2 exon 11 (NM_004972).
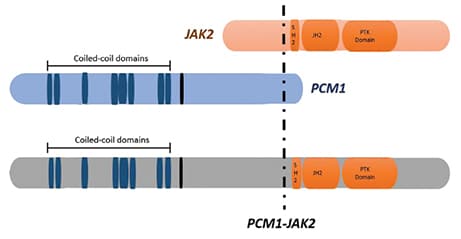
The PCM1 gene is located on the short arm of chromosome 8 and encodes a protein with multiple coiled-coil motifs that are essential for proper centrosome assembly. JAK2 is a tyrosine-protein kinase gene located on the short arm of chromosome 9. The fusion of PCM1–JAK2 can result from breakpoints at different exons of both PCM1 and JAK2. This chimeric gene product retains several of the coiled-coil motifs from PCM1 as well as the entire tyrosine kinase encoding domain of JAK2 resulting in constitutive activation of the JAK2 kinase domain (Fig. 3).5
Discussion. Molecular testing plays an important role in evaluating MPNs. Characteristic variants in JAK2, CALR, and MPL help confirm the diagnosis. However, this case highlights how coincidental variants in these genes can lead to premature diagnostic closure. The classic CALR exon 9 variants associated with MPNs are a 52 bp deletion (p.L367Tfs*46) and a 5 bp insertion (p.K385Tfs*47) resulting in a pathogenic frameshift in the CALR gene encoding the calreticulin protein.6,7 These mutations are considered mutually exclusive with JAK2 and MPL genes in MPNs, although rare cases of ET and PMF with concurrent JAK2 p.V617F and CALR mutations have been reported.8-11 CALR exon 9 mutations are not reported in PV. CALR exon 9 frameshift deletion/insertion has been associated with a favorable prognosis in MPNs as compared with JAK2 and MPL-mutated MPNs.7
The variant identified in this patient—CALR exon 9 deletion, c.1142_1144del—is a 3 bp in-frame deletion resulting in the deletion of glutamine (p.E381del). Interpretations of its significance have varied.6,12 It has been considered as an in-frame pathogenic germline mutation in a patient with a diagnosis of MPN-unclassifiable.6 In another study, this variant was identified in JAK2 p.V617F-positive ET patients by high-resolution melting curve analysis, but was wild type by Sanger sequencing.12 At the time of this analysis, the CALR c.1142_1144del has not been reported in somatic variant cancer databases, such as COSMIC and cBioPortal. In gnomAD, a database of genomic variants in healthy populations, this in-frame deletion has been reported with an overall allele frequency of 0.013 percent, with highest frequency reported in the Ashkenazi Jewish population (0.155 percent).13 While there is no definitive consensus, population frequencies above 0.1 percent are typically benign, representing normal genomic variation in the human population.14 Based on these data and the observed VAF of 50 percent in our case, this 3 bp in-frame deletion (p.E381del) of a single amino acid is likely a benign germline polymorphism. Studies have reported 3 bp, 9 bp, and 12 bp CALR exon 9 in-frame deletions, which are also considered likely germline polymorphisms.7,15-17 The presence of benign in-frame deletions, while rare, creates a potential diagnostic pitfall for molecular assays using polymerase chain reaction and fragment analysis. Since in-frame germline variants in exon 9 could be confused with pathogenic frameshift deletions by sizing methods, confirmation by MPS or Sanger sequencing is prudent when uncommon variants are detected.
In view of the detection of a PCM1–JAK2 rearrangement, the final diagnosis in this case was revised from CALR-positive myelofibrosis to PCM1–JAK2-fusion–positive myelofibrosis. Patients with PCM1–JAK2-fusion–positive neoplasms often present with hepatosplenomegaly, a leukoerythroblastic picture and neutrophilic precursors in the peripheral blood, and eosinophilia, although this feature may be absent in some cases. Only a mild increase in eosinophils was noted in the bone marrow aspirate smear for our patient. The marrow can also show fibrosis, similar to ET and PMF. Prognosis and survival are variable, and an aggressive clinical course with transformation to acute leukemia has been reported in the literature.4 Based on the diagnosis of a CALR-mutated MPN, a lower-risk MPN, the initial plan was to observe the patient with blood monitoring every three months without need for a systemic therapy. However, detection of PCM1–JAK2 rearrangement predicted higher-risk disease and changed her management to include a JAK2 inhibitor (ruxolitinib) as first-line therapy for symptom control. Because of a variable and nondurable response to tyrosine kinase inhibitors and a significant risk of disease progression,18 an allogeneic bone marrow transplant is planned as soon as possible for this patient.
Conclusion. As this case illustrates, the diagnostic workup of MPNs and related neoplasms requires the synthesis of complex clinical, pathologic, and laboratory data. Molecular diagnostics and interpretation of genomic variants is an important step in this process and may differ among laboratories based on adopted analysis strategies. Efforts are being made to standardize the approach to variant interpretation in cancer genomics.19 While there was clearly an initial error in the predicted protein consequence resulting from the CALR in-frame deletion described in this case, it serves as a reminder that coincidental, nonpathogenic germline variants do occur in disease-specific genes.20 Proper variant curation must rigorously apply objective data to classify variants and should avoid undue bias based on the patient’s diagnosis as this may lead to suboptimal patient care.
Variant curation is complicated, and having access to molecular pathology expertise can help improve patient care. Consult services are prevalent in multiple other areas of pathology and laboratory medicine; however, they are relatively new in the molecular genomic arena. Seeking a second molecular opinion, as illustrated in this case, can help identify discrepancies and recognize opportunities for additional high-yield testing.
- Swerdlow SH, Campo E, Harris NL, et al., eds. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. Rev. 4th ed. IARC Press; 2017:30–59.
- Kim SY, Im K, Park SN, Kwon J, Kim JA, Lee DS. CALR, JAK2, and MPL mutation profiles in patients with four different subtypes of myeloproliferative neoplasms: primary myelofibrosis, essential thrombocythemia, polycythemia vera, and myeloproliferative neoplasm, unclassifiable. Am J Clin Pathol. 2015;143(5):635–644.
- Bain BJ, Horny HP, Arber DA, Tefferi A, Hasserjian RP. Myeloid/lymphoid neoplasms with eosinophilia and rearrangement of PDGFRA, PDGFRB or FGFR1, or with PCM1–JAK2. In: Swerdlow SH, Campo E, Harris NL, et al., eds. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues. Rev. 4th ed. IARC Press; 2017:72–79.
- Reiter A, Gotlib J. Myeloid neoplasms with eosinophilia. Blood. 2017;129(6):704 –714.
- Reiter A, Walz C, Watmore A, et al. The t(8;9)(p22;p24) is a recurrent abnormality in chronic and acute leukemia that fuses PCM1 to JAK2. Cancer Res. 2005;65(7):2662–2667.
- Mikic TB, Pajic T, Sever M. CALR mutations in a cohort of JAK2 V617F negative patients with suspected myeloproliferative neoplasms. Sci Rep. 2019;9(1):19838.
- Klampfl T, Gisslinger H, Harutyunyan AS, et al. Somatic mutations of calreticulin in myeloproliferative neoplasms. N Engl J Med. 2013;369(25):2379–2390.
- Kang MG, Choi HW, Lee JH, et al. Coexistence of JAK2 and CALR mutations and their clinical implications in patients with essential thrombocythemia. Oncotarget. 2016;7(35):57036–57049.
- Cleyrat C, Chabot-Richards DS, Lynch DT, et al. Leukemic transformation of post-essential thrombocythemia myelofibrosis: a unique case presenting with double MPL and CALR mutations. Blood. 2017;130(suppl 1):4215.
- Jeromin S, Meggendorfer M, Fasan A, Haferlach C, Kern W, Haferlach T. Frequency of concurrent BCR-ABL1, JAK2, CALR and MPL mutations in a cohort of 5,545 cases with suspected MPN by a deep sequencing approach. Poster presented at: European Hematology Association Congress; June 22–25, 2017; Madrid, Spain. https://j.mp/2Y1LPT0.
- McGaffin G, Harper K, Stirling D, McLintock L. JAK2 V617F and CALR mutations are not mutually exclusive; findings from retrospective analysis of a small patient cohort. Br J Haematol. 2014;167(2):276–278.
- Lim KH, Chang YC, Gon-Shen Chen C, et al. Frequent CALR exon 9 alterations in JAK2 V617F-mutated essential thrombocythemia detected by high-resolution melting analysis. Blood Cancer J. 2015;5(3):e295.
- Genome Aggregation Database. V2.1.1. https://gnomad.broadinstitute.org/variant/19-13054604-AGAG-A?dataset=gnomad_r2_1. Accessed April 19, 2021.
- Sukhai MA, Misyura M, Thomas M, et al. Somatic tumor variant filtration strategies to optimize tumor-only molecular profiling using targeted next-generation sequencing panels. J Mol Diagn. 2019;21(2):261–273.
- Bilbao-Sieyro C, Florido Y, Gómez-Casares MT. CALR mutation characterization in myeloproliferative neoplasms. Oncotarget. 2016;7(33):52614–52617.
- Nangalia J, Massie CE, Baxter EJ, et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. N Engl J Med. 2013;369(25):2391–2405.
- Wang Y, Ho AK, Pan Q, Racke FK, Jones D. In-frame exon 9 CALR deletions co-occur with other alterations in the JAK-STAT pathway in myeloproliferative neoplasms. Blood. 2014;124(21):4588.
- Schwaab J, Naumann N, Luebke J, et al. Response to tyrosine kinase inhibitors in myeloid neoplasms associated with PCM1–JAK2, BCR–JAK2 and ETV6-ABL1 fusion genes. Am J Hematol. 2020;95(7):824–833.
- Li MM, Datto M, Duncavage EJ, et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a joint consensus recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn. 2017;19(1):4–23.
- Yang G, Viswanatha DS, He R. Frameshift and in-frame CALR exon 9 genetic alterations detected in a post-ET myelofibrosis patient before and after stem cell transplantation. AMP Case Report. CAP TODAY. 2020;34(2):28–32.
Dr. Gupta was a molecular genetic pathology fellow at time of writing, Dr. Frank is associate professor of clinical pathology and laboratory medicine, Dr. Roth is assistant professor of clinical pathology and laboratory medicine, and Dr. Priore is assistant professor of clinical pathology and laboratory medicine—all in the Department of Pathology and Laboratory Medicine, Hospital of the University of Pennsylvania Perelman School of Medicine, Philadelphia. Dr. Luger is professor of medicine, Department of Medicine, Division of Hematology and Oncology, Hospital of the University of Pennsylvania Perelman School of Medicine.
[divider]
Test yourself
Here are three questions taken from the case report.
1. What is the most common type of pathogenic CALR variant in myeloproliferative neoplasms?
a. Frameshift variant.
b. Missense variant.
c. Nonsense variant.
d. In-frame insertion or deletions.
2. The following statement is true about PCM1-JAK2–mutated myeloid neoplasms:
a. These neoplasms have favorable prognosis.
b. These neoplasms have a variable prognosis and require TKIs and allogeneic transplant in most cases.
c. These neoplasms can be observed and managed conservatively as compared with other myeloproliferative neoplasms.
d. Histopathological features are distinct from the more common myeloproliferative neoplasms.
3. Which of the following describes a frameshift variant?
a. CALR c.1142A>T.
b. CALR c.1142_1144del.
c. CALR c.1142_1143del.
d. CALR c.1142_1144dup.
Answers are online now at www.amp.org/casereports.

