Charna Albert
July 2019—Twenty years ago, Ulysses Balis, MD, bought the domain name HistoQuery.org. He had realized that one day, digital pathology would be sophisticated enough to be incorporated into interactive, Web-based tools, so he did what any self-styled geek would do with such a nascent idea—he waited.

“Histo” stands for histology, and “query” stands, unsurprisingly, for query, explains Dr. Balis, professor of pathology and director of the Division of Pathology Informatics at the University of Michigan. HistoQuery, he knew, would be the name of the free educational tool he would develop to draw on the collective expertise of the global pathology community. He just had to wait for the technology to catch up and think of a great website design.
That aha moment arrived at the 2019 USCAP meeting, in March. It was USCAP’s first year using an audience participation system so session attendees could be polled with multiple-choice questions during each presentation and respond via smartphone on the conference app. As Dr. Balis watched histograms paired with audience responses populate in real time on a presenter’s PowerPoint, he realized that with the right execution, a Web-based version of the same technology could be used to pose difficult cases and, in turn, receive votes on the most likely diagnosis from a community of pathology experts—all without the need for page refreshes.
After the conference, Dr. Balis brought the idea to his department. In less than two months, the information technology team had developed a working version of HistoQuery.org, which was demoed at the 2019 Pathology Informatics Summit, in May.
Much like the audience participation system that inspired the site, HistoQuery is a crowdsourcing tool. After a brief registration process, users can post a new case or vote on responses to other cases. They can also provide a new diagnosis, or theory, as HistoQuery terms them, along with the rationale behind that theory. (Users can only submit one theory, and while they can change their vote, they can’t vote twice.) While the team hasn’t developed a standalone mobile app, the website employs responsive Web architecture designed to accommodate and adjust to desktop and mobile device-based Web browsers.
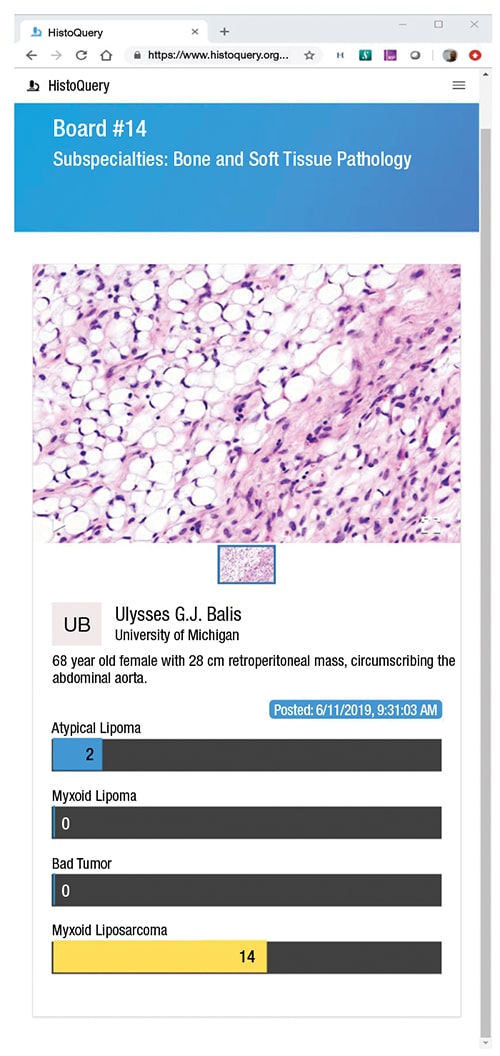
The cases, or boards as they’re called on HistoQuery, are typically composed of a slide image or images and a case history. The case histories on the site are brief. Two examples: “68 year old female with 28 cm retroperitoneal mass, circumscribing the abdominal aorta” (see screenshot above) and “Nasal mass at 1×, 10×, and 40×. What is your diagnosis and what + and − stains would support it?”
By responding to questions posed on the site, Dr. Balis wrote in a May post on the Association for Pathology Informatics listserv, “you become an essential component of the crowdsourced equation.” The tool, he added in his post, could be helpful for “seeking consultation from your colleagues on a particularly difficult case in a weekly slide teaching set. Simply post an image or two along with any salient history . . . and . . . wait for the flow of answers and their associated histogram to appear, in real time.”
Before HistoQuery, Dr. Balis says, there was no real solution for “when you’re crunched for time going through cases and need an immediate answer.” Pathologists had begun to use social media to share digital pathology images from rare cases, and some software vendors provide clearinghouses for posing questions about cases to other users. But what sets HistoQuery apart from these other offerings is that it facilitates real-time crowdsourcing within a free, Web-based global forum.
“I think this tool could benefit trainees by providing a safe and anonymous environment for posing questions about cases,” says Scott Owens, MD, gastrointestinal and hepatobiliary pathologist and director of the Division of Quality and Health Improvement at the University of Michigan. “Everybody who’s been through a residency recognizes there are questions about a case you’d rather ask your peers than an attending.”

Dr. Owens, a colleague of Dr. Balis who was not involved in developing HistoQuery, further explains that the tool has the potential to improve upon the practice of pathology trainees sharing information about cases on social media sites, which aren’t organized to retrospectively analyze how a consensus was reached on a diagnosis. HistoQuery “fills an educational gap in that it provides a central forum and repository for sharing cases,” he says.
The bar graphs on the HistoQuery site look like elements of a static Web page, similar to figures in an online medical journal. Behind the scenes, though, each response contains what’s known as a Web sockets interface. This interface, which was built with a library of programming conventions called ReactJS, allows images rendered on a Web page to update in real time according to changes made to a database. Put another way, each histogram is linked to a database row that updates every time a new vote is cast, allowing the results to tabulate in real time without requiring page refreshes.
“The minute the database is updated, another table links the updated row in the database to every browser that is currently logged on to the HistoQuery site, triggering each bar graph to update to a new value,” Dr. Balis says. The back end of the system is a Postgres database housed at the University of Michigan.
Refreshing a browser now and then, a regular feature of much online activity, might not seem to be unduly limiting. However, Dr. Balis is looking ahead to mass global adoption of the site. “If 500 users were logged in at the same time,” he notes, “you can imagine the value of the graphs updating very quickly.” His team has done testing with thousands of simulated users on the system, proving the site can accommodate a critical mass of concurrent transactions without lagging.
“The thing that will make or break HistoQuery isn’t the technology—that part is ready,” Dr. Balis says. Less certain is the human element: Will it achieve the level of adoption needed to become a true global forum for pathology information sharing?
At CAP TODAY press time, the site had about 250 registered users. Dr. Balis acknowledges that it’s going to take a while to get the word out, but he’s optimistic that upcoming improvements to HistoQuery will drive adoption and keep users engaged. These include the following:
- Personalized notifications: During registration, users are prompted to choose pathology subspecialties of interest. Eventually, they will be given the option to receive an email notification when a new case, or a board, is posted in their chosen subspecialties.
- A geospatial map displaying the location of people logged in to the site. (“It’s always satisfying to know where you’re helping people,” Dr. Balis says.)
- A rolling comment board for each case, a chat-like feature Dr. Balis hopes will foster discussion.
- An element of gamification in the form of a leader board announcing which users have posted the highest number of responses to cases. (“Friendly competition tends to drive adoption in other crowdsourcing Web applications,” he notes.)
In addition, the team has near-term plans to incorporate a high-performance whole slide viewer developed by the Balis Lab for the Kidney Precision Medicine Project, an initiative of the National Institute of Diabetes and Digestive and Kidney Diseases. And it plans to add Viper Studio, a Balis Lab open-source application that generates ground truth maps of foreground regions of interest—in other words, a tool that can create data for training machine-learning algorithms to recognize salient features of histology images.
Consequently, Dr. Balis is optimistic that HistoQuery, if widely adopted, could become the “entry point to a larger scale opportunity involving computation and machine learning.” Theoretically, crowdsourcing tools such as HistoQuery have the potential to generate statistics about the population of a given rare disease entity much more quickly and easily than other methodologies. Therefore, if the global community adopts HistoQuery en masse, Dr. Balis says, the tool could become “a publicly curated repository of rare cases” that could be used by machine-learning experts to train algorithms to assist pathologists in rendering suggested diagnoses for rare cases.
“The most difficult part of machine-learning activities,” Dr. Balis continues, “is generating the curated datasets needed for training, particularly with rare cases. The beauty of this crowdsourcing vehicle is you get rare cases in tandem with the crowdsourced wisdom of the most likely diagnosis. It’s everything you need to create a curated training repository.”

Bruce Friedman, MD, emeritus professor of pathology at the University of Michigan, agrees. “As the creater of HistoQuery,” Dr. Friedman says, “Dr. Balis doesn’t want to be in a position to speculate about how the tool might be used in the future [for other than training purposes]. I’m not encumbered by any of those same restrictions.”
Dr. Friedman, who also was not involved in developing HistoQuery, is interested in how the tool can be used by practicing pathologists. If the site attracts a sufficient amount of traffic, he says, surgical pathologists could use crowdsourcing in a very practical way to buttress convictions about a particular case in their daily work queue. “A practicing pathologist would never use this tool for a challenging case where he or she didn’t already have a good idea of the diagnosis or for which they required a consultation by a national expert,” he continues. Therefore, he’s not suggesting that HistoQuery would replace the services of consulting pathologists. However, it could provide diagnostic support in cases where a surgical pathologist is already fairly certain of the diagnosis.
Dr. Friedman too imagines the tool could have global health implications, particularly in developing countries with an acute shortage of pathology services. By using HistoQuery, he says, clinicians in those countries would, at the very least, receive a crowdsourced diagnosis from the international community.
“Dr. Balis is appropriately calling this a teaching tool, for now,” Dr. Friedman says. And in this capacity, he adds, HistoQuery has the potential to provide rich material for pathology training programs in smaller hospitals. “Someone who trains in pathology at a prestigious institution will come across a broad array of unusual cases,” he explains. “As part of the curriculum, it could be stipulated that residents at [smaller hospitals] review cases submitted to HistoQuery on a continuing basis, which, in turn, would expose them to a broader array of cases.”
In every subspecialty, says Dr. Owens, picking up the thread of the conversation, there are tricky diagnoses that even the experts routinely have trouble with—in gastrointestinal pathology, a good example is the grading of dysplasias in precancerous conditions of the esophagus, such as Barrett’s esophagus. “We see thousands of those cases here [at the University of Michigan],” he explains, “but there are places where pathologists don’t have that opportunity.” HistoQuery, he adds, could expose trainees and pathologists to a greater array of such cases.
To this end, Dr. Owens conducted a test of the site’s usefulness by submitting a deidentified image from his caseload. He had already secured a consult on the image, a case of fungal infection in the GI tract, from a colleague who sub-subspecializes in such contagions. While some of the answers submitted to HistoQuery differed from what he knew to be the correct diagnosis, the accurate response received the majority of the votes.
Dr. Owens foresees situations in which pathology departments use HistoQuery for intramural consultations as well, particularly institutions that have buildings throughout a city, with experts in different subspecialties located blocks or miles from each other. In that situation, HistoQuery could be used to poll various specialists on a case more quickly and easily than via email.
Taking its potential use a step further, Drs. Owens and Friedman concur that HistoQuery could be integrated into a laboratory’s quality assurance practices. Laboratories might “upload one in 10 cases to HistoQuery” and receive a “sufficient number of crowdsourced diagnoses to function as a QA tool,” Dr. Friedman says.
Dr. Balis is quick to acknowledge that it was the programming team in Michigan’s Division of Pathology Informatics that allowed such possibilities for HistoQuery. The site, he explains, uses much of the same code as PathTrack, the real-time geospatial pathology specimen tracking system the University of Michigan implemented last summer. (See “For one laboratory, a workflow transformation,” CAP TODAY, June 2018, www.captodayonline.com.) “The good news is [the team] was already primed on how to build an ultra-high concurrency application that works in real time. Building this tiny little application was child’s play compared to PathTrack,” he says with a laugh.
HistoQuery, he adds, could not have been completed so quickly if Michigan’s Department of Pathology didn’t have its own full-stack IT development team—evidence he uses to make the case that pathology departments should maintain at least some degree of independent programming capability. “Great ideas come along in every department. If you’re entirely dependent on central IT and you tell them you want to finish a project in eight weeks, they’re going to look at you like you’ve got three eyes,” he says, half-joking.
Dr. Balis sought to develop and release HistoQuery in a short time frame because he envisioned it fulfilling an urgent and fundamental need. “It crossed my mind that if the tool could be implemented correctly, you would have one of the so-called killer apps for pathology education which needs to exist but doesn’t.”
Adds Dr. Friedman, “He’s the expert in this. If he calls it a killer app, I’ll call it a killer app.”
Charna Albert is a writer in Chicago.

