Summary
CAP TODAY and the Association for Molecular Pathology have teamed up to bring molecular case reports to CAP TODAY readers. AMP members write the reports using clinical cases from their own practices that show molecular testing’s important role in diagnosis, prognosis, and treatment. The following report comes from City of Hope Comprehensive Cancer Center, Duarte, Calif., and Cedars-Sinai Medical Center, Los Angeles. If you would like to submit a case report, please send an email to the AMP at amp@amp.org. For more information about the AMP and all previously published case reports, visit www.amp.org.
Parastou Tizro, MD
Eric Vail, MD; Ronald Paquette, MD
Celeste Eno, PhD, D(ABMGG)
December 2025—Acute promyelocytic leukemia with PML::RARA is a unique subtype of acute myeloid leukemia with specific biology and clinical characteristics. APL makes up five to eight percent of AML cases in younger patients but is less common in the elderly.1 It is characterized by predominance of abnormal promyelocytes and presence of a hallmark translocation t(15;17)(q24;q21), resulting in fusion of the promyelocytic leukemia (PML) gene with the retinoic acid receptor alpha (RARA) gene.2,3
Clinically, this fusion provides an increased sensitivity to specific therapeutic agents, such as all-trans-retinoic acid (ATRA) in combination with chemotherapy or arsenic trioxide (ATO). This treatment has changed APL from a fatal to a highly curable leukemia (approximately 90 percent cure rate), but rapid and accurate diagnosis is crucial to start life-saving therapy promptly.4 Although PML::RARA is found in up to 95 percent of APL cases with typical morphology, it is worth noting that atypical isoforms and variant RARA translocations also have been described.4,5 The various functional domains of the PML protein retained within the PML::RARA atypical isoforms or different RARA partners could potentially result in different clinical outcomes and response to therapy.4,6
Here we describe a PML::RARA rearrangement with an atypical breakpoint in PML. This case highlights the significance of multiple testing techniques and inability to use common measurable residual disease testing platforms to follow in remission, owing to potentially false-negative results.
Case. A 21-year-old male with a reported history of APL in remission for three years after receiving chemotherapy with ATRA plus ATO presented to the clinic with progressive shortness of breath and pallor. Laboratory evaluation revealed severe pancytopenia. His blood smear showed rare circulating promyelocytes. Bone marrow biopsy revealed predominantly granulocytic precursors consisting almost entirely of abnormal promyelocytes, with virtually no mature forms. Erythroid and megakaryocytic precursors were suppressed by leukemia but showed no overt dysplasia. A differential count showed 85 percent promyelocytes. Flow cytometry demonstrated 82 percent blast equivalents with expression of CD13, CD33, CD38, CD64, CD117, and MPO, and lacking CD34 and HLA-DR. A preliminary diagnosis of AML suggestive of APL was rendered, and the patient was started on ATRA pending confirmation of diagnosis. He achieved morphologic remission with a combination of ATO and ATRA.
A blood sample was sent for PML::RARA quantitative reverse transcriptase polymerase chain reaction (qRT-PCR) because of concern for the relapsed leukemia. The result was very low-level positive for short isoform (bcr3) at 0.02 percent. qRT-PCR was repeated on the bone marrow and showed only low-level positivity for PML::RARA fusion short form (bcr3) at 0.09 percent. A rapid AML next-generation sequencing panel by amplicon capture, which covers RARA fusions with 17 different partners including PML, detected fusion reads at the limit of detection. Fluorescent in situ hybridization using a dual-color dual-fusion probe set was positive for PML::RARA rearrangement in 179/200 (89.5 percent) of the cells scored on bone marrow aspirate. Due to this discrepancy, Anchored Multiplex PCR (AMP) next-generation sequencing was performed, which enables the detection of novel fusion partners and breakpoints. The results confirmed the presence of an atypical bcr2 transcript involving a novel breakpoint within exon 6 of PML and a breakpoint in intron 2 of the RARA gene (Fig. 1).
Discussion. Based on current guidelines, both morphologic and immunophenotypical studies and confirmatory genetic analysis are part of the diagnostic workup of APL. The confirmatory tests may include cytogenetic analysis for t(15;17) by karyotype and/or FISH and molecular testing by qRT-PCR methods for PML::RARA.2-4,7 Although PML::RARA is found in up to 98 percent of APL cases with typical morphology, rare instances may evade detection by standard qRT-PCR.5,8
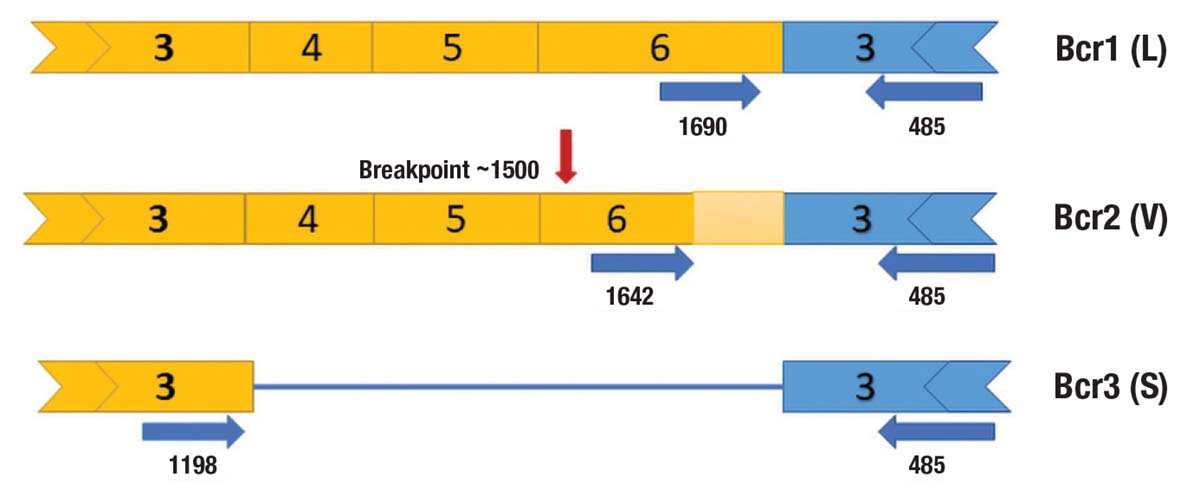
Three distinct typical PML::RARA transcript isoforms can be discerned based on specific breakpoints within the PML gene: 1) the long isoform (intron 6, known as bcr1); 2) the variant isoform (exon 6, known as bcr2); and 3) the short isoform (intron 3, known as bcr3).5,9 Only one breakpoint in intron 2 has been identified in the RARA gene in these cases.5,10
Rare atypical PML::RARA transcripts have been reported, with the most common being atypical bcr2 (V-forms). These arise from mis-splicing or breaks involving a novel donor site in PML exon 6 and are typically unique.11,12 These splicing events in PML can be linked to concomitant insertions of nucleotides from RARA intron 2.12,13 Moreover, partial deletions of RARA exon 3 and alternative isoforms of RARA have been observed in atypical transcripts.14-16
The majority of reported APL cases with atypical isoforms can be detected using routine qRT-PCR methods; however, in a few cases, as in our case, this assay may not be able to be used to establish a diagnosis.12 Specifically, the primer for bcr2 is located downstream of the patient’s breakpoint (Fig. 1), and the bcr3 primer was able to detect trace signals due to the immense amount of product at diagnosis. However, bcr3 primer would not be expected to identify product during treatment and clinical remission. Therefore, knowledge of these atypical cases is essential for monitoring measurable residual disease.17 False-negative results would have serious implications for patient therapy, as early treatment intervention in patients displaying MRD positivity leads to more favorable outcomes.4 FISH can be used as a disease surveillance tool in this case; however, this assay is not sensitive enough for effective MRD monitoring. Similar to qRT-PCR, amplicon-based NGS platforms may fail to detect variant isoforms due to the primer design. Although AMP NGS is a potential alternative NGS method for detecting atypical isoforms, this technology is not capable of detecting transcripts at MRD. A patient-specific qRT-PCR assay could be designed based on the sequence derived from AMP NGS for personalized MRD monitoring.
Patients with classical PML::RARA fusions typically respond well to retinoid therapy and have a favorable prognosis. However, this may not hold true for atypical or variant isoforms, which can be associated with variable outcomes and treatment responses. Therefore, beyond confirming the presence of a PML::RARA fusion, precise identification of the transcript is crucial for appropriate clinical management and for accurately interpreting outcomes in retinoid-based clinical trials.4,9
In summary, we report a case in which routine qRT-PCR and amplicon-based sequencing failed to detect the atypical PML::RARA fusion isoform, which was later identified by AMP NGS. Although the patient achieved hematologic remission, the lack of a suitable method for molecular monitoring limits effective follow-up. This highlights the need for agnostic or universal qRT-PCR primers capable of detecting PML::RARA fusions regardless of breakpoint location.
- Chen Y, Kantarjian H, Wang H, Cortes J, Ravandi F. Acute promyelocytic leukemia: a population-based study on incidence and survival in the United States, 1975-2008. Cancer. 2012;118(23):5811–5818.
- de Thé H, Chomienne C, Lanotte M, Degos L, Dejean A. The t(15;17) translocation of acute promyelocytic leukaemia fuses the retinoic acid receptor α gene to a novel transcribed locus. Nature. 1990;347(6293):558–561.
- Borrow J, Goddard AD, Sheer D, Solomon E. Molecular analysis of acute promyelocytic leukemia breakpoint cluster region on chromosome 17. Science. 1990;249(4976):1577–1580.
- Sanz MA, Fenaux P, Tallman MS, et al. Management of acute promyelocytic leukemia: updated recommendations from an expert panel of the European LeukemiaNet. Blood. 2019;133(15):1630–1643.
- Liquori A, Ibañez M, Sargas C, Sanz MA, Barragán E, Cervera J. Acute promyelocytic leukemia: a constellation of molecular events around a single PML-RARA fusion gene. Cancers (Basel). 2020;12(3):624.
- Huang W, Sun GL, Li XS, et al. Acute promyelocytic leukemia: clinical relevance of two major PML-RARα isoforms and detection of minimal residual disease by retrotranscriptase/polymerase chain reaction to predict relapse. Blood. 1993;82(4):1264–1269.
- Beillard E, Pallisgaard N, van der Velden VHJ, et al. Evaluation of candidate control genes for diagnosis and residual disease detection in leukemic patients using ‘real-time’ quantitative reverse-transcriptase polymerase chain reaction (RQ-PCR)—a Europe against cancer program. Leukemia. 2003;17(12):2474–2486.
- Geoffroy MC, de Thé H. Classic and variants APLs, as viewed from a therapy response. Cancers (Basel). 2020;12(4):967.
- Grimwade D, Biondi A, Mozziconacci MJ, et al. Characterization of acute promyelocytic leukemia cases lacking the classic t(15;17): results of the European Working Party. Blood. 2000;96(4):1297–1308.
- Pandolfi PP, Alcalay M, Fagioli M, et al. Genomic variability and alternative splicing generate multiple PML/RAR alpha transcripts that encode aberrant PML proteins and PML/RAR alpha isoforms in acute promyelocytic leukaemia. EMBO J. 1992;11(4):1397–1407.
- Gallagher RE, Li YP, Rao S, et al. Characterization of acute promyelocytic leukemia cases with PML-RARα break/fusion sites in PML exon 6: identification of a subgroup with decreased in vitro responsiveness to all-trans retinoic acid. Blood. 1995;86(4):1540–1547.
- Iaccarino L, Divona M, Ottone T, et al. Identification and monitoring of atypical PML/RARA fusion transcripts in acute promyelocytic leukemia. Genes Chromosomes Cancer. 2019;58(1):60–65.
- Ismail S, Ababneh N, Awidi A. Identification of atypical PML-RARA breakpoint in a patient with acute promyelocytic leukemia. Acta Haematol. 2007;118(3):183–187.
- Cao Y, Yao L, Liu Y, et al. An atypical PML-RARA rearrangement resulting from submicroscopic insertion of the RARA gene at the PML locus with novel breakpoints within PML exon 7b and RARA exon 3. Acta Haematol. 2019;142(2):98–104.
- Jezíšková I, Rázga F, Gazdová J, et al. A case of a novel PML/RARA short fusion transcript with truncated transcription variant 2 of the RARA gene. Mol Diagn Ther. 2010;14(2):113–117.
- Park TS, Kim JS, Song J, et al. Acute promyelocytic leukemia with insertion of PML exon 7a and partial deletion of exon 3 of RARA: a novel variant transcript related to aggressive course and not detected with real-time polymerase chain reaction analysis. Cancer Genet Cytogenet. 2009;188(2):103–107.
- Ramos Perez JM, Patel KP, Loghavi S, et al. Value of measurable residual disease monitoring in patients with acute promyelocytic leukemia in the era of frontline ‘chemotherapy-free’ therapy. Leuk Lymphoma. 2022;63(3):672–675.
Dr. Tizro is assistant professor of hematopathology and molecular genetics, Department of Pathology and Laboratory Medicine, City of Hope Comprehensive Cancer Center, Duarte, Calif. Dr. Vail is assistant professor of pathology and laboratory medicine, Cedars-Sinai Medical Center, Los Angeles. Dr. Paquette is professor of medicine, director of myeloid malignancies, and clinical director of the stem cell and bone marrow transplant program, Department of Hematology and Oncology, Cedars-Sinai Medical Center. Dr. Eno is assistant professor of pathology and laboratory medicine and associate director of the molecular pathology laboratory, Cedars-Sinai Medical Center.

