Editor: Frederick L. Kiechle, MD, PhD
Submit your pathology-related question for reply by appropriate medical consultants. CAP TODAY will make every effort to answer all relevant questions. However, those questions that are not of general interest may not receive a reply. For your question to be considered, you must include your name and address; this information will be omitted if your question is published in CAP TODAY.
[button color=”blue” size=”small” link=”https://captodayonline.com/q-a-submission/” target=”blank” ]Submit a Question[/button]
Q. Is there a place for procalcitonin testing alongside PCR testing on the BioFire system? Our facility has saved a tremendous amount of money on decreased lengths of stay by implementing a sepsis protocol that uses the typical emergency department treatment as well as lactic acid levels and PCR, which gives us a definitive response in about an hour after a blood culture turns positive. Is there any way that PCT can be seen as an enhancement to this process?
A. March 2020—It appears that this laboratory is using the BioFire blood culture identification test and is curious whether procalcitonin would add value. If so, my answer would be a definitive maybe. There are at least two areas in which PCT might add value. The first is related to typical ED treatment, as mentioned. In many places, this treatment would be an initial dose of broad-spectrum antibiotics while the blood culture/BioFire results are pending. With appropriate turnaround time and clinician buy-in, PCT results might help reduce this initial treatment. The caveat is that while the literature shows strong negative predictive value for PCT, most clinicians with whom I have dealt are hesitant to withhold that first dose based on PCT. There is a strong desire to do something while awaiting the culture/BioFire result.
Still, getting that initial PCT would be important for subsequent measurements and dose de-escalation. The BioFire system is not going to cover all antimicrobial resistance mechanisms. It hits the Klebsiella pneumoniae carbapenemase but doesn’t cover imipenemase, New Delhi metallo-β-lactamase, or oxacillinases, for instance, with regard to carbapenemases. In addition, these PCR assays will struggle with novel variants that are only detectable phenotypically (a small percent but still a consideration). These challenges mean that PCT could still have a role in guiding antimicrobial stewardship.
All that said, adoption and trust of the BioFire results are much easier to achieve than adoption and trust of PCT, in my experience and opinion. It would take substantial buy-in and trust from your clinical colleagues to make sure the expense of PCT leads to improved patient outcomes.
Joshua Hayden, PhD, DABCC
Chief of Chemistry
Norton Healthcare
Louisville, Ky.
A. The topic of PCT-enhancing molecular diagnostics is interesting. PCT can help optimize the need for certain molecular tests as it does with antimicrobial use and traditional microbiology testing. Unfortunately, there have not been large-scale studies combining PCT with molecular pathogen detection, but clearly there is value in helping to differentiate bacterial versus nonbacterial etiologies and helping to decide which molecular panels should be used. We are a bit constrained technologically at this time since molecular infectious disease tests are often run as panels (with the exception of newer singleplex/two-plex point-of-care devices). Many of these panels include viral and bacterial pathogens, which negates the benefit of PCT for this role. However, random-access molecular platforms where you can pick and choose pathogens by individual species, groups, or clusters will be available in the future and could better leverage PCT.
For nonrespiratory infection molecular testing, PCT (with other clinical parameters) may help improve pretest probability—that is, patients who would benefit from additional testing. An example is the new prediction rule by investigators at the University of California, Davis, for febrile infants (Kuppermann N, et al. JAMA Pediatr. 2019;173[4]:342–351). The rule included PCT and other variables (urinalysis and absolute neutrophil count) and can rule out more serious issues, such as meningitis, and perhaps make it possible to avoid a lumbar puncture (and not require use of a molecular meningitis panel). In the end, as PCR panels expand in terms of detectable pathogens and flexibility, and perhaps with the adoption of other novel technologies beyond PCR, PCT can help optimize utilization and add to the predictive power of these tests. On our end, we’ve seen PCT-negative results (along with other clinical factors like a normal lactate, for example) help reduce unnecessary cultures and antimicrobial therapy as reported by other studies. Integrating PCT with other workflows that include molecular diagnostics is the next logical step.
Nam K. Tran, PhD, HCLD(ABB)
Associate Clinical Professor
Director of Clinical Chemistry,
Special Chemistry and Toxicology, Point-of-Care Testing, and SARC
Department of Pathology and Laboratory Medicine
UC Davis School of Medicine, Davis, Calif.
Q. We are looking into validation methods for lower breakpoints for carbapenems. The CDC has banks of organisms available to run on our Vitek 2 (BioMérieux). Are there guidelines on which organisms to include and how many times to test?
A. This question is particularly pertinent because the Clinical and Laboratory Standards Institute updated more clinical breakpoints in 2019 than it has in any single year since 2010.1 Keeping up to date with antimicrobial susceptibility test breakpoints is a challenging but critically important task for the clinical laboratory that impacts the quality of patient care and patient safety.
Many laboratories struggle with the breakpoint validation process. In particular, there is uncertainty as to why a validation study must be performed at all since the analytical output—that is, the minimal inhibitory concentration—is the same, and only the interpretation is changed when implementing a new breakpoint. However, it is useful to reflect on the performance characteristics required by the FDA for AST system clearance. These include essential agreement (MIC agreement within ±1 doubling dilution) and categorical agreement (susceptible [S], intermediate [I], and resistant [R] interpretation agreement with a reference method). Systems may be tweaked by the manufacturer to ensure that the performance around a given breakpoint is ideal and both factors can be met. This tweaking may result in unacceptable performance at lower or higher MICs not bracketing the breakpoints. When the breakpoint is changed, the adjustments made thereafter may no longer be appropriate, and the FDA categorical agreement requirements may not be met. Another reason labs should validate the updated breakpoints is that the bacteria may have changed, due to changing and evolving resistance mechanisms, between the time the labs’ commercial systems were first FDA cleared and the time breakpoints were updated by the CLSI and/or FDA. For example, systems that are applying outdated carbapenem breakpoints have not been updated since 2009—a time when CREs were rare in the U.S. Therefore, laboratories should want to ensure these systems perform as well as they did when such resistance mechanisms were rare. In both cases, the laboratory should be aware of how the test system performs with the new breakpoints.
When it comes to performing a validation study, several resources are available to the clinical laboratory. CLSI publishes the M52 guideline, “Verification of Commercial Microbial Identification and Antimicrobial Susceptibility Testing Systems,” which includes, in appendix B, a section on how to perform a validation for breakpoint updates.2 Similarly, the CLSI AST News Update published an issue that reviewed breakpoint validation.3 Another excellent resource is the materials generated by Janet Hindler that are available on the California Department of Health’s website.4 This resource includes step-by-step instructions, worksheets, and troubleshooting instructions for how to perform a validation study for carbapenems and Enterobacterales. The tools can be adapted to other breakpoint validations as well.
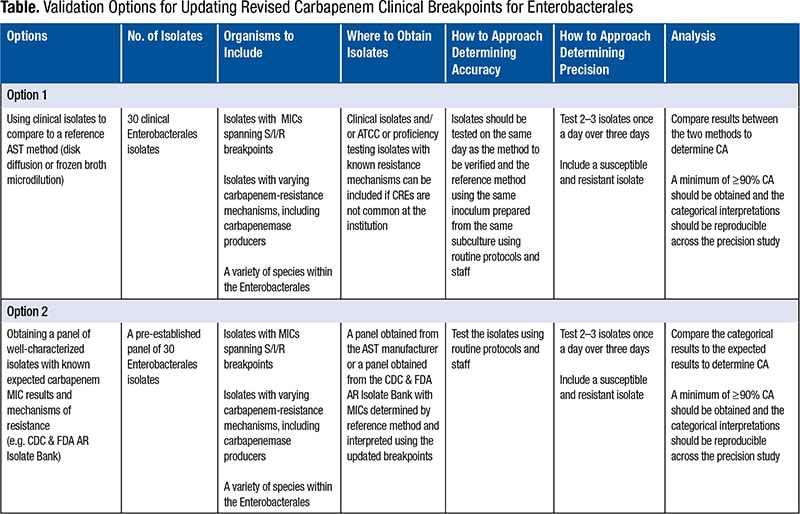
In general, a validation study for a breakpoint update should be minimal as compared with a study performed when implementing a brand-new AST system in the laboratory. In both cases, a study verifies (as the name implies) the performance characteristic established by the manufacturer. For breakpoints that have not yet been FDA cleared on a system, the manufacturer cannot provide performance data to the laboratory, so the laboratory should establish, in advance of the study, what acceptable performance looks like and validate that its test can achieve that performance. In most cases, this is greater than 89 percent categorical agreement, although laboratories may need to be flexible, at the laboratory director’s discretion, for some drug/bug combinations if performance falls below this. While typical validation studies include evaluations of accuracy (agreement) and precision (reproducibility) of essential agreement and categorical agreement, essential agreement testing is not needed since it was determined for the dilution range on the panel when the laboratory first implemented the test. The Table describes two options for updating the revised carbapenem clinical breakpoints for Enterobacterales. Accuracy testing across 30 isolates with a variety of S/I/R results is generally viewed as sufficient, with comparison to a reference or known result. Ideally, the lab would also test a variety of species within the Enterobacterales and include a variety of resistance mechanisms, such as carbapenem-resistant isolates that do and do not produce carbapenemases, when verifying the carbapenem breakpoints. The Centers for Disease Control and Prevention and FDA Antibiotic Resistance Isolate Bank is an excellent resource for such isolates since the isolates are well characterized and available at no charge.5 Even more useful is a specific panel (the CDC’s Enterobacteriaceae Carbapenem Breakpoint Panel) designed to validate the carbapenem breakpoints for Enterobacterales, which takes the guesswork out of which isolates to choose for the validation study. The CDC isolates are sent to the laboratory with a known reference MIC and interpretation by current breakpoints. Validation of accuracy can, therefore, be as simple as testing the CDC isolates on the lab’s AST system and comparing the interpretation (with current breakpoints) from that system with the interpretation provided by the CDC.
Precision can be accomplished by testing two to three isolates once a day (intra-assay reproducibility) over three days (inter-assay reproducibility) and ensuring the interpretation is the same across the replicate tests. Precision validation can use clinical isolates, CDC isolates, isolates obtained from proficiency testing challenges, or even QC strains.
It is important to note that neither the CAP nor the Clinical Laboratory Improvement Amendments dictate the extent of studies required to validate or verify any test. Rather, the testing protocol should be determined by the laboratory’s medical director and be scientifically justifiable. Nonetheless, the guidelines listed herein are generally accepted by the clinical community as sufficient to ensure that AST system performance is acceptable with the current breakpoints.
- Humphries RM, Abbott AN, Hindler JA. Understanding and addressing CLSI breakpoint revisions: a primer for clinical laboratories. J Clin Microbiol. 2019;57(6):e00203-19.
- Clinical and Laboratory Standards Institute. M52: Verification of Commercial Microbial Identification and Antimicrobial Susceptibility Testing Systems; 1st ed.; 2015.
- Do you need help with verification of your AST system? CLSI AST News Update [newsletter]. Wayne, Pa: Clinical and Laboratory Standards Institute; December 2016. https://clsi.org/media/1700/clsi-news-winter-2016.pdf.
- Healthcare-associated infections (HAI) program. California Department of Public Health website. https://www.cdph.ca.gov/Programs/CHCQ/HAI/Pages/CA_ARLN.aspx. Updated Dec. 20, 2019. Accessed October 2019.
- CDC and FDA Antibiotic Resistance (AR) Isolate Bank. Centers for Disease Control and Prevention website. https://www.cdc.gov/drugresistance/resistance-bank/index.html. Reviewed Feb. 14, 2020. Accessed October 2019.
Additional references
Bartsch SM, Huang SS, Wong KF, et al. Impact of delays between Clinical and Laboratory Standards Institute and Food and Drug Administration revisions of interpretive criteria for carbapenem-resistant Enterobacteriaceae. J Clin Microbiol. 2016;54(11):2757–2762.
Clinical and Laboratory Standards Institute. CLSI supplement M100: Performance Standards for Antimicrobial Susceptibility Testing. 29th ed.; 2019.
Clinical and Laboratory Standards Institute. M23: Development of In Vitro Susceptibility Testing Criteria and Quality Control Parameters. 5th ed.; 2018.
FDA-recognized antimicrobial susceptibility test interpretive criteria. U.S. Food and Drug Administration website. https://www.fda.gov/drugs/devel opment-resources/fda-recognized-antimicrobial-susceptibility-test-interpretive-criteria. Current as of Aug. 27, 2019. Accessed October 2019.
Humphries RM, Ferraro MJ, Hindler JA. Impact of 21st Century Cures Act on breakpoints and commercial antimicrobial susceptibility test systems: progress and pitfalls. J Clin Microbiol. 2018;56(5):e00139-18.
Romney M. Humphries, PhD, D(ABMM), M(ASCP)CM, MT(ASCP)
Chief Scientific Officer, Accelerate Diagnostics
Professor, Department of Pathology
University of Arizona College of Medicine
Tucson, Ariz.
Member, CAP Microbiology Committee
Patricia Jeanne Simner, PhD, D(ABMM)
Associate Professor of Pathology, Johns Hopkins Medicine
Baltimore, Md.
Member, CAP Microbiology Committee

